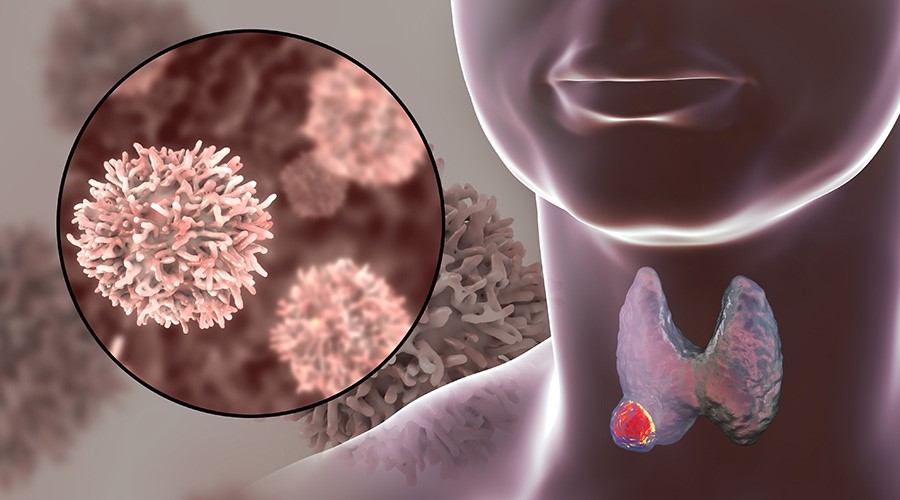
Malignant transformation is a process that involves genetic alterations, such as point mutations and chromosomal abnormalities, and epigenetic alterations, such as DNA methylation, histone post-translational modifications, and promoter hypermethylation. All of these variations influence cell growth, apoptosis, and differentiation. In more advanced stages of carcinogenesis, other changes may promote angiogenesis, invasion of adjacent tissues, and metastasis to distant sites. The best-known genes related to cancer code for proteins and are classified as oncogenes and tumour suppressor genes.
However, one of the great surprises of modern biology was the discovery that only about 2% of the genes that make up the human genome code for proteins. In recent years, with the introduction of high-throughput techniques for studying gene expressions, such as microarrays and whole transcriptome sequencing, it has been determined that at least 90% of the genome is actively transcribed and that the transcriptome human is more complex than the set of genes that encode proteins since it exhibits a significant expression of non-coding RNAs.Among them are microRNAs (miRNAs), which participate in gene regulation mechanisms, and their altered face has been shown to play an essential role in the malignant transformation of human cells.
CHARACTERISTICS OF miRNAs
miRNAs are small RNA molecules (approximately 22 nucleotides) that are conserved through evolution and are involved in post-transcriptional gene silencing. They are present in multicellular organisms and viruses. They are located on all human chromosomes except the Y chromosome. About 50% are clustered, and transcription is polycistronic. They are frequently situated in fragile sites and regions of amplification or loss of heterozygosity associated with cancer. They may be located in intergenic areas or introns of protein-coding genes; less frequently, they reside in exons but have an antisense orientation relative to the protein-coding gene.
It is estimated that 1-5% of the human genome corresponds to miRNAs, which can regulate at least 30% of the genes that encode proteins. Each miRNA can interact with hundreds of messenger RNAs (mRNAs) either directly or indirectly. Similarly, the expression of a single mRNA can be cooperatively modulated by multiple miRNAs.
BIOGENESIS AND PROCESSING OF miRNAs
The biogenesis of miRNAs begins in the nucleus with the transcription of the corresponding genes by RNA polymerase II or III to give rise to a primary miRNA (pri-miRNA), which is polyadenylated and has a 7-methyl-group attached. Guanylate (m7G) at the 5′ end, which is known as capping. Subsequently, this primary transcript is processed by the enzyme Drosha to form a precursor miRNA (pre-miRNA), about 70 nucleotides long, that resembles a hairpin. This is transported to the cytoplasm by exportin-5 (XPO5), where the Dicer enzyme processes it into a 22-nucleotide duplex miRNA. One of the strands of the duplex interacts with the RNA silencing-induced complex (RISC) and, in this way, interacts with target mRNA while the complementary chain is degraded. There are two mechanisms of negative regulation of mRNAs by miRNAs. Whether one or the other occurs is governed by the degree of complementarity between the miRNA and the target mRNA. If this is imperfect, the translation of the mRNA is repressed. The complementarity sites for the miRNAs that use this mechanism are located in the non-translated regions, towards the 3′ end of the mRNA (3’UTR areas). If, on the other hand, the complementarity is perfect, a cut is induced, and subsequent degradation of the mRNA occurs. The miRNAs that use this mechanism find their complementarity sites in the coding sequences of the mRNAs.
miRNA FUNCTIONS
More than 1,000 miRNAs have been identified in humans through cloning experiments and bioinformatics. They regulate numerous metabolic and cellular pathways, notably those that control modifications during development, embryogenesis, stem cell preservation, hematopoietic cell differentiation, and brain development. Altered expression of miRNAs is likely to contribute to human disease and, among other processes, has been linked to tumour progression, including tumour growth, differentiation, adhesion, apoptosis, invasion, and invasion—metastasis formation.
The first reports on miRNAs were in the worm Caenorhabditis elegans, where investigations led to the description of genes that code for small RNAs related to developmental changes in this species. Until then, it was believed that it was an exclusive phenomenon in nematodes, but a series of works were carried out that allowed the identification and cloning of miRNAs from different organisms, including humans. It was discovered that the nucleotide sequences were phylogenetically conserved.
The first evidence linking miRNAs to cancer comes from a study in patients with chronic lymphocytic leukaemia (CLL), which examined a recurrent deletion located on chromosome 13q14.3. The smallest common region of the deletion was found to code for two miRNAs: miR-15a and miR-16-1, suggesting their role as tumour suppressor genes. When these miRNAs are expressed commonly, they bind to the 3’UTR region of the mRNA of the anti-apoptotic protein BCL2, which causes the inhibition of its translation, and the usual mechanisms of programmed cell death can be activated. The absence of miR-15a and miR-16-1 induces high levels of this protein and the blocking of apoptosis. Other examples of miRNAs that function as tumour suppressors are the families of let-7 and miR-34.
miRNAs can also act as oncogenes. The best-studied example is the miR-17-92 cluster. This includes six mature miRNAs (miR-17, miR-18a, miR-19a, miR19b-1, miR20a, and miR-92-1) that share a standard primary transcript generated from the 13q31.3 loci. The cluster is amplified in various lymphomas and lung, colon, pancreas and prostate cancer. Its expression can be directly regulated by the transcription factors c-Myc and E2F. Overexpression of this cluster is associated with tumour development. Similarly, miR 21, miR155 and miR272/miR273 are other examples of miRNAs that act as oncogenes.
In short, when the expression of miRNAs is altered, their gain or loss of function is triggered in cancer cells, which is why the definitions of oncogenes and tumour suppressors have expanded to include miRNAs, in addition to the classic genes that encode proteins. Another aspect of interest is that the expression patterns of miRNAs are tissue-specific so that the same miRNA can act as an oncogene or a tumour suppressor, depending on the context.
MECHANISMS THAT ALTER THE EXPRESSION OF miRNAs IN HUMAN CANCER
The altered expression of miRNAs is the primary mechanism that triggers their gain or loss of function in cancer cells. The activation of oncogenic transcription factors such as myc is another important mechanism that alters the expression of miRNAs. Another pathway may be the occurrence of chromosomal aberrations. Increased expression of miRNAs has been associated with genomic amplification, and decreased expression has been associated with chromosomal deletion, in addition to other mechanisms such as point mutations and mutations. Aberrant methylation of promoters. On the other hand, global repression of miRNA biogenesis emerges as a cancer-specific mechanism, as mutations in critical components of the miRNA processing machinery, such as Drosha, DICER1, and XPO5, promote malignant transformation and carcinogenesis.
CLINICAL APPLICATIONS OF miRNAs
Since the altered expression of miRNAs is related to cancer development and metastasis formation, they have great potential to function as biomarkers for disease status and progression, as well as for diagnosis, prognosis, classification and evaluation of risk factors. In this sense, they present some advantages, such as that mature miRNAs are relatively stable, the study of their expression does not require large amounts of samples, and they can be measured in biopsies of fresh tissue. They have even been detected in tissue fixed in formalin and embedded in paraffin. Recent studies show that they can also be measured in some biological fluids such as serum/plasma or saliva, which offers a less invasive way of screening. miRNA expression profiles have been used to distinguish tumour samples from normal tissues, to identify tumour tissue of unknown origin or poorly differentiated tumours, as well as to distinguish different tumour subtypes. Some alterations of miRNAs occur in patients at early stages, so that they can be helpful for the early detection of cancer.
From a predictive point of view, its usefulness has been demonstrated as an indicator of clinical outcome, 55 of the tendency to recurrence and metastasis. Additionally, it can be a predictor of response to a given treatment. miRNAs have not only been detected in cancerous tissue but also in surrounding tissue, so they can be used to detect alterations in the tumour microenvironment. Single nucleotide polymorphisms (SNPs) within genes encoding miRNAs or their molecular targets are suspected to be detrimental and may increase an individual’s risk of developing diseases such as cancer.
Some strategies have been explored for therapeutic purposes to normalise the expression of miRNAs. One of them has the objective of reducing the expression of miRNAs with oncogenic action. To this end, modified anti-miRNAs (OMA) oligonucleotides have been synthesized, known as “antagomirs”, which are complementary to endogenous miRNAs and allow their specific inhibition. For its application in the clinic, it will be necessary to achieve its effective release in the target tissue, an aspect that is under investigation. Similarly, a new type of miRNA inhibitors called “miRNA sponges” have been developed, which contain multiple sites to bind target miRNAs and inhibit them with the same efficacy as OMAs. Another strategy consists of raising the expression of miRNAs with tumour suppressor function. This can be achieved using liposomes, polymers, nanoparticles or viral vectors that contain miRNAs with reduced expression and thus restore their normal levels. These novel designs require further evaluation to become therapeutic opportunities for cancer patients.
FUTURE PERSPECTIVES
Research on miRNAs should focus on the identification of tissue-specific molecular signatures that regulate metastasis; explore the miRNAs that play an essential role in the regulation of cancer stem cells; to translate the advances of the laboratory into the development of new prognostic markers and new therapeutic strategies, as well as to develop new techniques for the detection of miRNAs.